Regression discontinuity with sci-kit learn models
from sklearn.gaussian_process import GaussianProcessRegressor
from sklearn.gaussian_process.kernels import ExpSineSquared, WhiteKernel
from sklearn.linear_model import LinearRegression
import causalpy as cp
WARNING (pytensor.tensor.blas): Using NumPy C-API based implementation for BLAS functions.
%config InlineBackend.figure_format = 'retina'
Load data
data = cp.load_data("rd")
data.head()
| x | y | treated | |
|---|---|---|---|
| 0 | -0.932739 | -0.091919 | False |
| 1 | -0.930778 | -0.382663 | False |
| 2 | -0.929110 | -0.181786 | False |
| 3 | -0.907419 | -0.288245 | False |
| 4 | -0.882469 | -0.420811 | False |
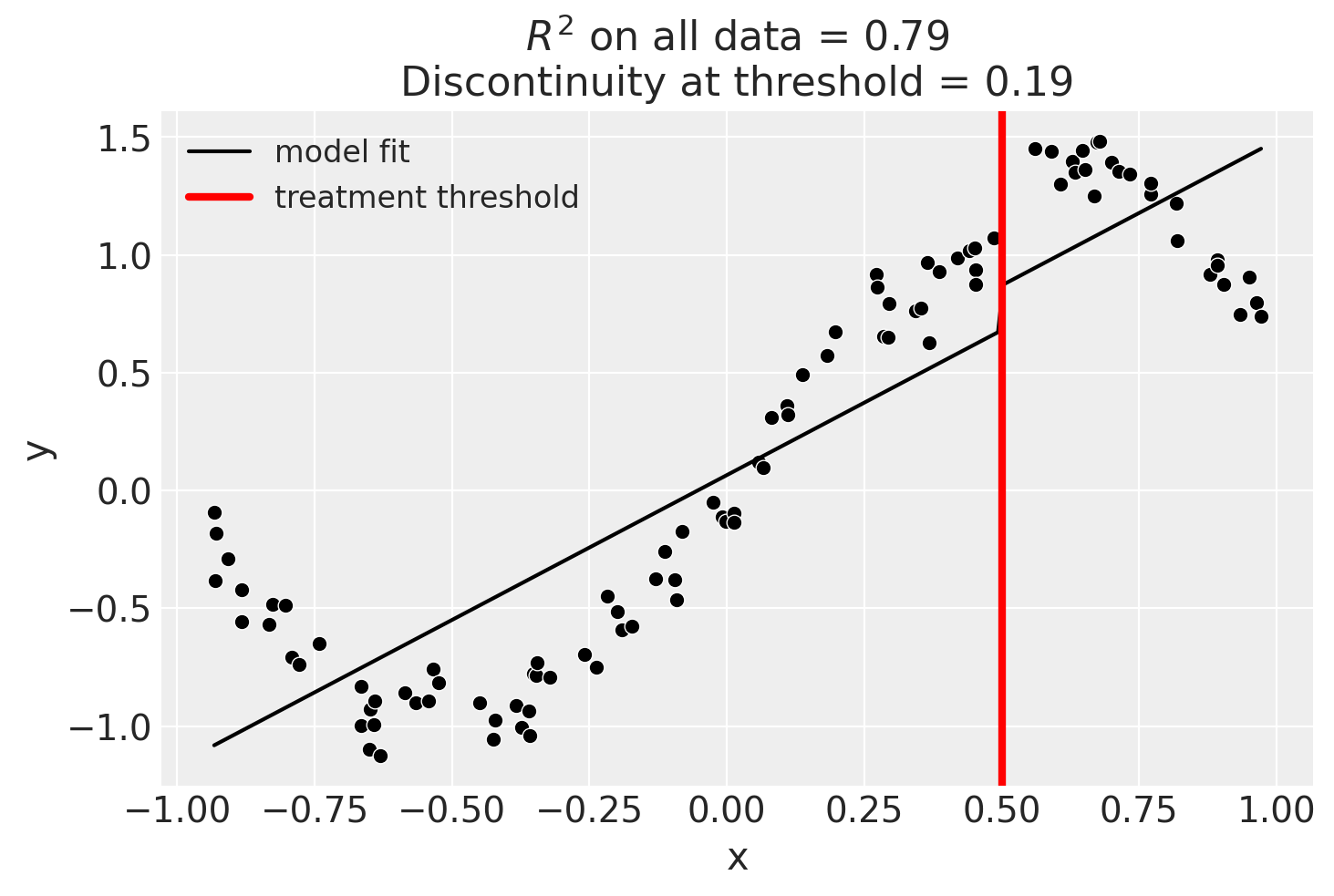
Linear, main-effects model
result = cp.skl_experiments.RegressionDiscontinuity(
data,
formula="y ~ 1 + x + treated",
model=LinearRegression(),
treatment_threshold=0.5,
)
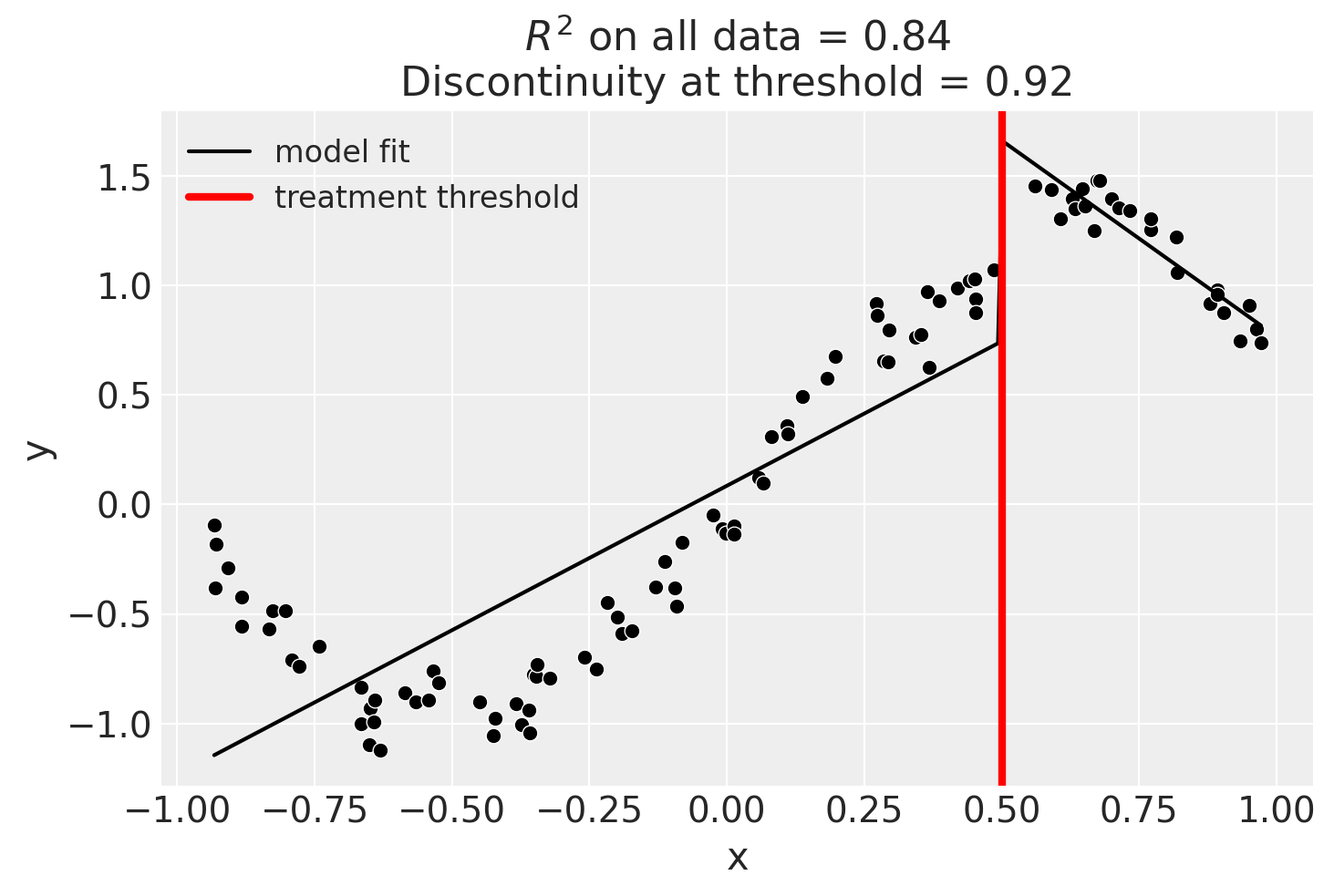
Linear, main-effects, and interaction model
result = cp.skl_experiments.RegressionDiscontinuity(
data,
formula="y ~ 1 + x + treated + x:treated",
model=LinearRegression(),
treatment_threshold=0.5,
)
Though we can see that this does not give a good fit of the data almost certainly overestimates the discontinuity at threshold.
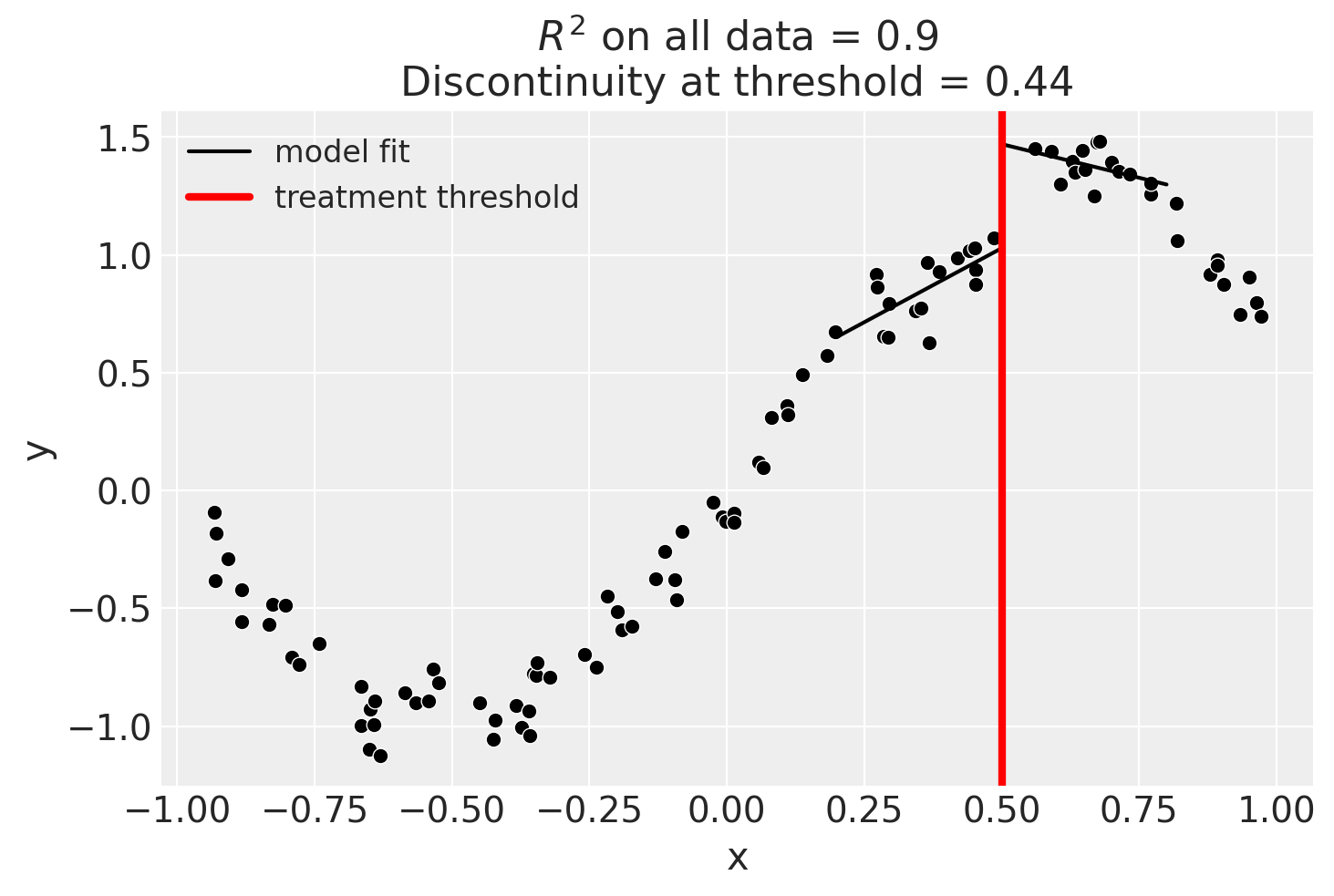
Using a bandwidth
One way how we could deal with this is to use the bandwidth kwarg. This will only fit the model to data within a certain bandwidth of the threshold. If \(x\) is the running variable, then the model will only be fitted to data where \(threshold - bandwidth \le x \le threshold + bandwidth\).
result = cp.skl_experiments.RegressionDiscontinuity(
data,
formula="y ~ 1 + x + treated + x:treated",
model=LinearRegression(),
treatment_threshold=0.5,
bandwidth=0.3,
)
result.plot();
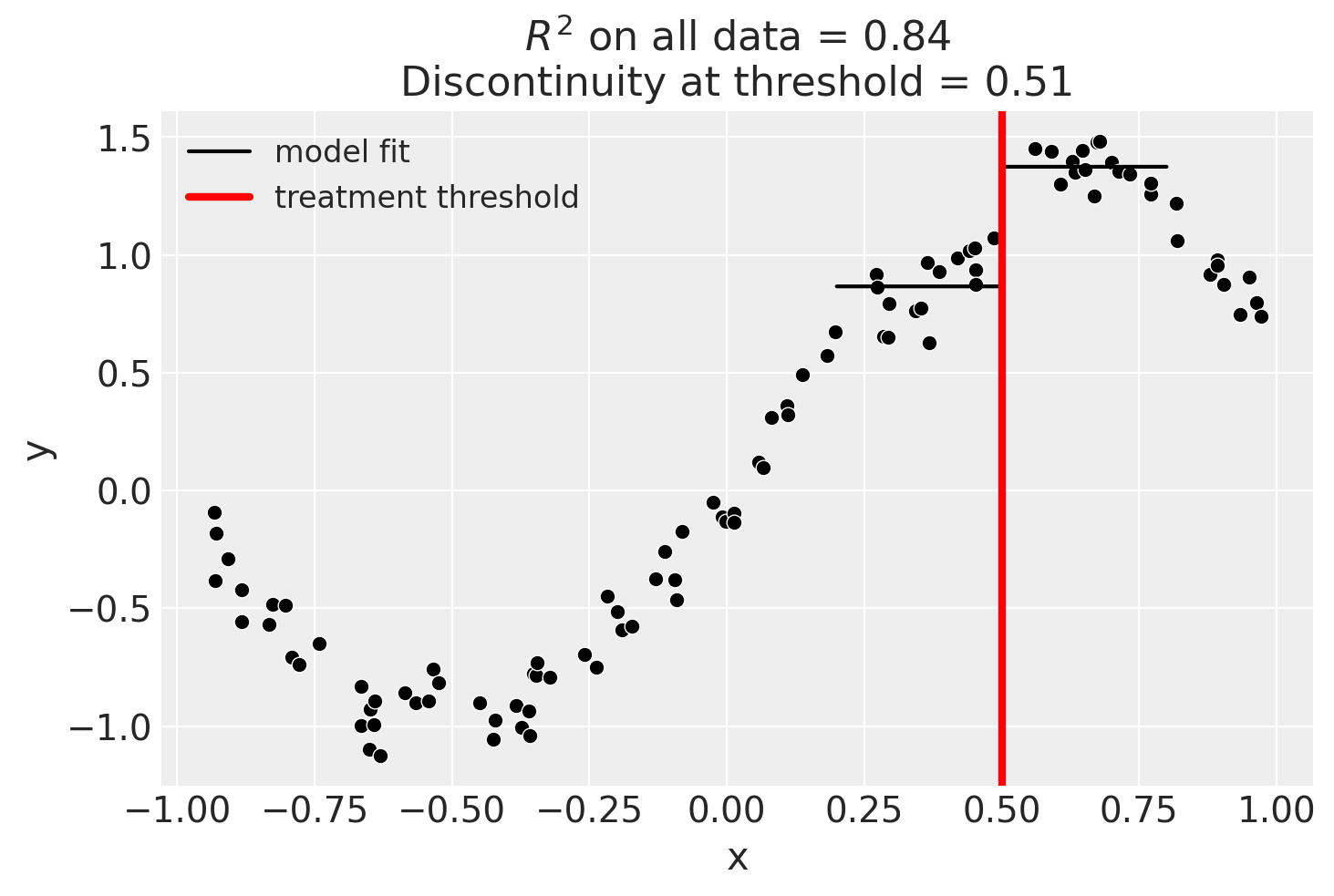
We could even go crazy and just fit intercepts for the data close to the threshold. But clearly this will involve more estimation error as we are using less data.
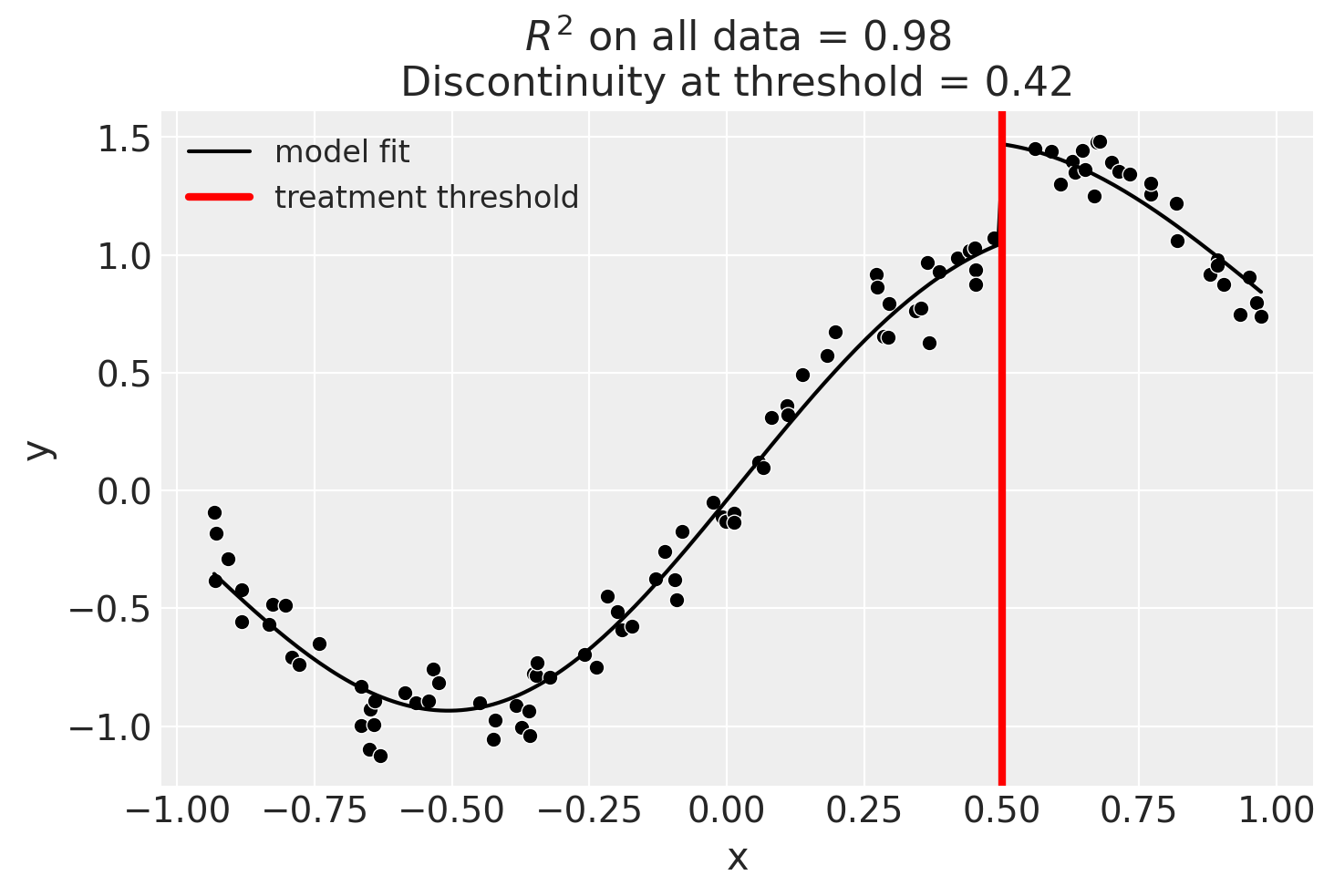
Using Gaussian Processes
kernel = 1.0 * ExpSineSquared(1.0, 5.0) + WhiteKernel(1e-1)
result = cp.skl_experiments.RegressionDiscontinuity(
data,
formula="y ~ 1 + x + treated",
model=GaussianProcessRegressor(kernel=kernel),
treatment_threshold=0.5,
)
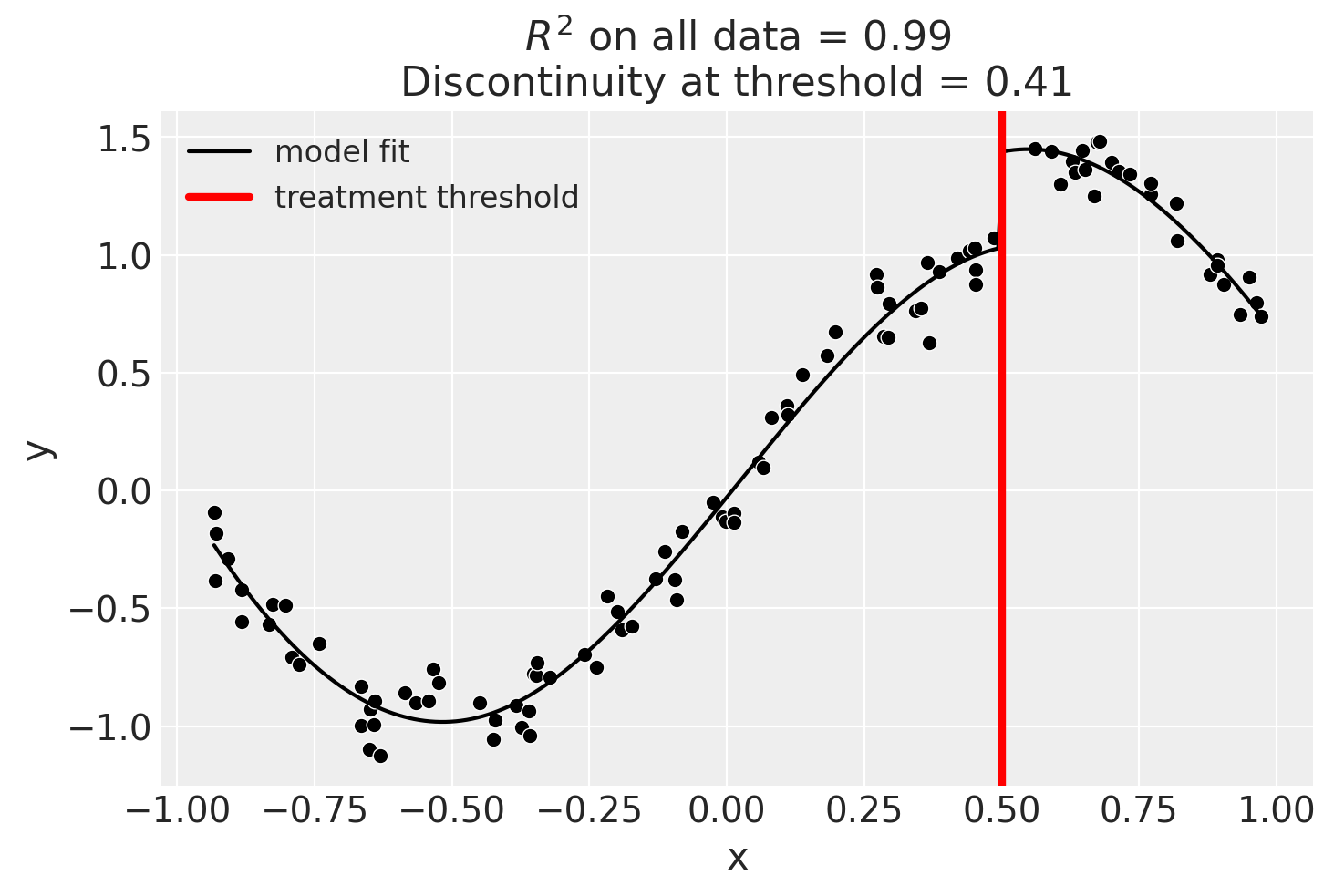
Using basis splines
result = cp.skl_experiments.RegressionDiscontinuity(
data,
formula="y ~ 1 + bs(x, df=6) + treated",
model=LinearRegression(),
treatment_threshold=0.5,
)